The Challenge
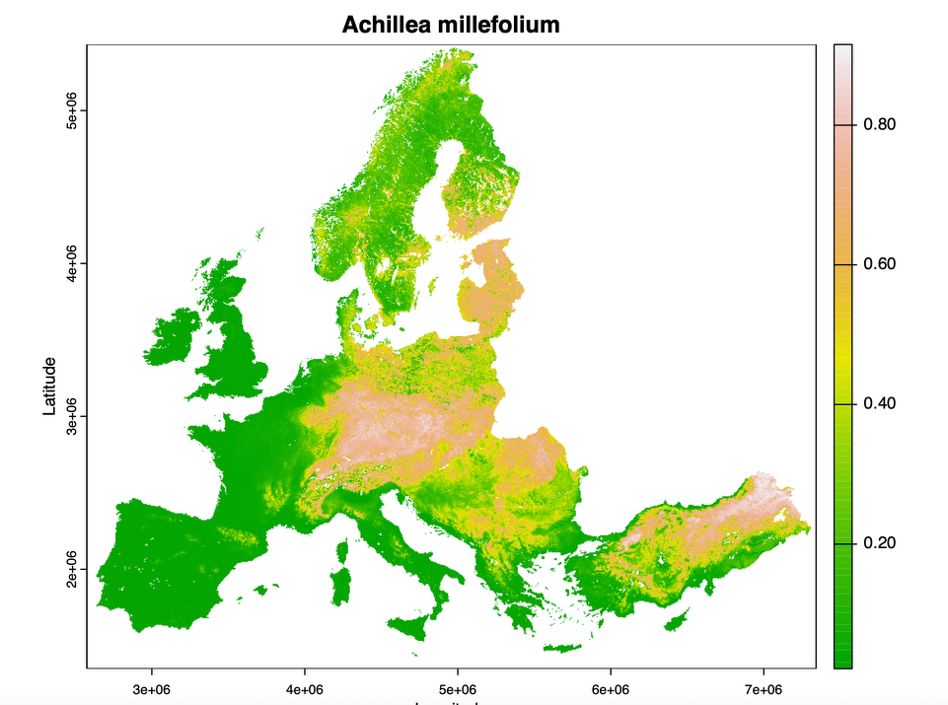
BioScore3.0 is a spatial explicit species specific biodiversity modelling framework used to assess the impacts of policy measures on biodiversity in Europe. BioScore uses relationships between the occurrence of species and environmental conditions to predict the effect of policy measures on the distribution of species and vegetation types across Europe.
BioScore3.0 used for the WET Horizon project which required the framework to grow in scope and application. Thus, there was a need to improve its structure, reproducibility, and capacity to incorporate additional taxonomic groups and enhance its transparency and collaborative potential.
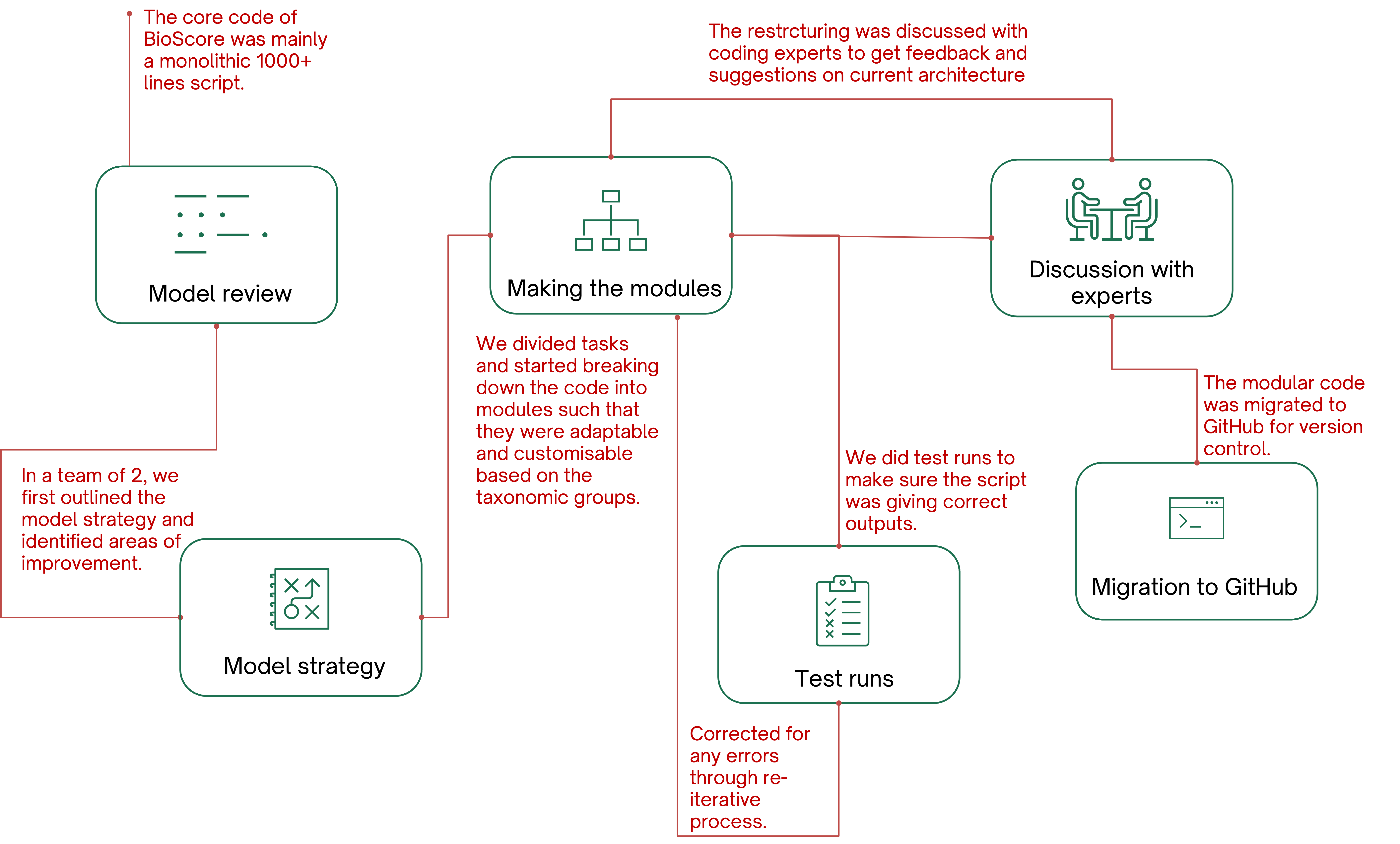
I collaborated on redesigning the workflow, incorporating new taxonomic groups, and implementing version control and collaborative practices using Git and GitHub. I initiated the BioScore project meeting for development of model strategy in line with the PBL's model directive.